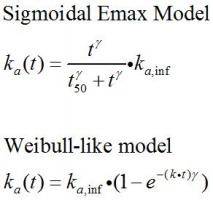Q: Can V be lower than total volume of blood and if yes, what is the meaning? if not what are the reasons?
A: V is the apparent volume of distribution and represents the degree to which the drug distributes into the tissues. Note that it is not a volume in the usual sense of a pool of liquid. Rather it is a value that enables us to scale between the administered drug (mass) and measured concentration (mass/volume).
Q: Can you please elaborate reason for selection of intravenous rather than extravascular for zero order input model?
A: This was answered during the demo. Selecting Extravascular in the built-in model forces the drug input through an absorption compartment with a 1st order rate constant Ka, and adds the differential equation deriv(Aa = -Ka*Aa). For the 0 order input, we are modeling a constant input, that is not proportional to the amount, to the central compartment with a specified duration, Tabs.
Q: Do you always have a fixef and stparm statement for each parameter?
A: Great question. The PML code that is written by the built-in options always specifies both a structural (stparm) and fixed effect (fixef) for each model parameter. This is to support population modeling, so that the structural parameter statements can be modified with random effects. The fixef statement then allows one to enter initial estimates and boundaries for the fixed effect. For individual modeling, there are no random effects, so it is not necessary to have both statements - only the fixef statement is required. The model with only fixed effects is as follows:
fixef(V = c(, 100, ))
fixef(Cl = c(, 45, ))
fixef(Tabs = c(, 4, ))
Even in population mode, if you have one parameter that has only a fixed effect you do not need to define the stparm but fixef is enough
Q: For Zero order input, Is it possible to select "infusion possible" option and then use A1 Rate =dose/Tabs?
A: Yes, this was also discussed in the demonstration. The zero order input can be entered as a rate, e.g. A1Rate = A1Dose/Tabs, instead of as an amount and duration.
Infusion possible is an option that is linked to the data. It automatically create an rate column and the program is calculating the duration for you as it knows that A1Rate=dose/duration .
Q: How does the model fitting performance change if you selected a different error model? other instead of additive error model?
This particular data set has similar results for both an additive and multiplicative error model. But for the additive error model, the final solution is somewhat sensitive to the initial estimate of CEps. Note that the CEps is an actual parameter in the model to be estimated and one should use an appropriate initial estimate for this parameter just like you would for any structural model parameter (V, Cl). It is generally a good strategy to use an initial estimate that is a bit higher than what you think the final estimate may be. Experience has taught us this often improves convergence.
Q: How would you add a Tlag to the 0-order model?
A: This can be done in the graphical or textual mode.
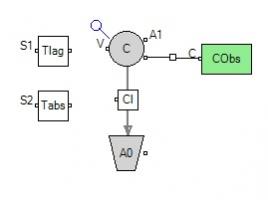
In Graphical mode, add both Tlag and Tabs as parameters as shown below:
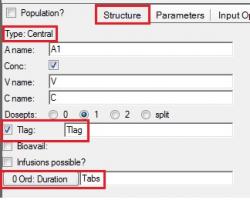
Then, click the Central compartment, and define Tlag and Tabs as inputs on the Structure tab:
In the Text mode, both Tlag and Tabs can be added to the dosepoint statement:
dosepoint(A1, tlag = (Tlag), duration = (Tabs))
Q: V, Tlag, Tabs were quite different in the slide shown after the demo and in the Phoenix demo. Could you comment?
A: This particular data set has similar results for both an additive and multiplicative error model. But for the additive error model, the final solution is somewhat sensitive to the initial estimate of CEps. Note that the CEps is an actual parameter in the model to be estimated and one should use an appropriate initial estimate for this parameter just like you would for any structural model parameter (V, Cl). It is generally a good strategy to use an initial estimate that is a bit higher than what you think the final estimate may be. Experience has taught us this often improves convergence.The screenshot was taken with the default additive error model term, CEps, set to 1. This is then fitted as a parameter under MLE. When the initial estimate for CEps is set to 10, the model fit and parameter precision improve.
Q: Can you use infusion possible with contant infusion?
A: Yes – you would then need to enter the amount (A1) and the rate (A1 Rate). An alternative is to use amount and duration (Tabs) as shown in the demo.
Q: Could we have a variable Ka depending on the part of the intestine? and if yes, how could we model that?
A: it you want to include a sophisticated absorption (PBPK based) it would be better to use a program specific for absorption modeling. But here are some other options. If you can also collect IV data then you could do numerical deconvolution to get a better estimate of the Rx release rate over time. In addition one could always fit quasi-mechanistic models for Ka. Transit models can be easily fit as can models such as:
Q: When setting initial estimates graphically bounds were not considered. How would you incorporate bounds?
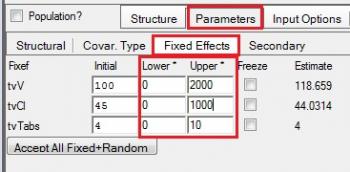
A: The initial estimates and boundaries can be set on the Parameters Tab, Fixed Effects sub-tab. Please see the screenshot below which incorporates bounds.
In general we recommend not applying bounds because the minimization algorithms prefer to explore values between negative infinity and positive infinity. However, if you need to use them, it is best to use one of lower or upper bounds, and not both. A better option would be to use mathematical transformations that prevent values from reaching impossible values. For example, if a parameter is exponentiated, it can never be less than zero. Mathematical transformations exist to bound values in nearly every way possible. Please note, QRPEM does not use boundaries even if you write down lower and upper bounds.
Q: I would like to clarify my question. As was shown earlier using single dose plasma data, can multiple dose plasma data be used to estimate the parameters? Not simulation using single dose model
A: Yes, although only simulation was shown in the demo, it is a similar setup for fitting, you need to specify the dosings (times and amounts) and then you can run a simple fit. There are examples of fitting models to multiple dose input data in the Gabrielsson and Weiner reference: PK4, PK11.
Q: What is A1 rate?
A: This is input for an infusion rate, this is added by default but not needed in this case‑
Q: If there were multiple profiles/patients, can you simulate each using their individual PK estimates?
A: Yes, this was shown in the Q&A during the demonstration. The input dataset must have a subject ID/patient column which is used as a Sort in the model. One can then go to Setup | Parameters, check the internal worksheet checkbox, and enter initial PK parameter estimates for each subject.
Q: Can you please show us how to model and simulate zero order input with first order absorption?
A. You would need to add zero order input and first order absorption each as dosepoint statements. This approach was discussed in the webinar PK43: multiple absorption routes.
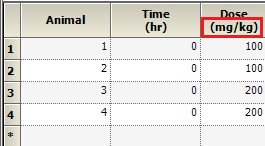
Q: If you have dose information as mg/kg only for animals, how to input for Aa?
A: It is possible to enter your dose amount in mg/kg, however it is important to understand that the output parameters will also be scaled by bodyweight in this case ( V in L/kg, Cl in L/hr/kg). To get units for your output parameters, we recommend using a Dosing input sheet with units as shown below. Then, map this in as the dosing sheet under Setup | Dosing.
Q: Is it possible to share the Phoenix file of rawdata and modeling?
A: No, we are not allowed to share the project file, however we will make the textual mode code available. This code can be easily imported into a Phoenix model object via cut-and-paste. Once imported you can use your own data to be fitted to the model or you can run simulations.