Hi all,
I have a cross-over study, each subject dosed three time (three food states) and each time his laboratory data was remeasured. Basically at this stage I want to use the base model to diagnose if any covariate change affects the parameter.
As I didn't select anything and want to diagnose from the eta-covariate graph, it only reports one eta-covariate for each ID. If I select the connection (say HDL with Cl), then it assumes the effects exist and I can't recheck from the eta-covariate graph any more. So I wonder at this stage how can I diagnose the potential covariates from a large pool (many combinations)???
Please notice that I have tried IOV and reset option, but as I mentioned above, I can only check the box to compare one at a time, rather than check all the eta-covariate graph (which can be done is non cross-over study and each ID has one set of covariates) , as it only reports one set of parameter for each ID rather than one?
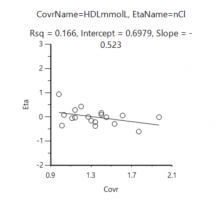
The fig below shows a typical eta-covariate graph, which contains all IDs, but only connect with the HDL in the first dose.
I also want to ask in a general way, if a covariate (biomaker) changes with a parameter (clearance) as the time varies, possible to report the Cl at each time point so a eta-covariate can be ploted? Or I can only try this through trial and error and compare the optimization each time?