Dear all,
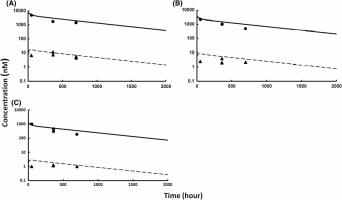
I’m working on brain modeling and penetration prediction, and I would like to use Dr Shah’s model. It’s a full PBPK model with a brain compartment divided in different sub-spaces. So I code it on Phoenix. In a first step, I wanted to validate the human version of the model with plasma data. So based on a reference described in the study I created the dataset for the model.
I have 2 questions:
- The profile changes based on #Points used in the simulation. For the same X range (Max X), comparing 100 vs 1000 #points, the simulated points (DV) are different for a same time (IVAR). From my understanding of Phoenix software, I expected to find the same value for a same time point. Could you explain me why we observed different simulated point for a same time point? What does the term “#points” mean exactly?
- Similarly, the difference between profiles from “Ind DV,IPRED vs IVAR” plot and “Ind Simulation” plot is not clear in terms of modeling. Even if the predictions in the first plot are not well, it follows the trend of the observations. Whereas in the “Ind Simulation” plot, the profile is fully different (when #points = 1000). From my understanding, this should lead to close results.
The project file is enclosed to this post in order to allow you to a look into it. The model is quite big but not that complex.
Many thanks for your help,
Béatrice
 ForumSim.phxproj 1.16MB
307 downloads
ForumSim.phxproj 1.16MB
307 downloads